Ancillary Generation Pipeline#
The generation of an ancillary file is usually carried out by one or more Ancillary-file-science scripts, as run by one or more tasks within Ancillary Workflows.
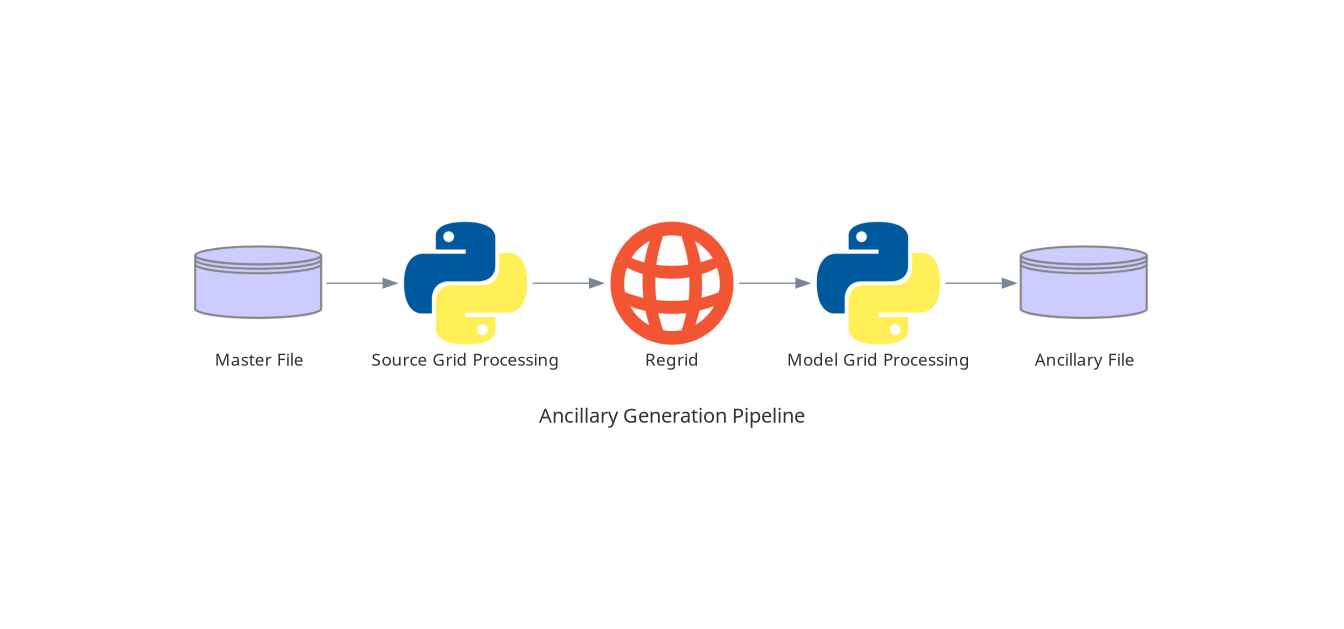
Regardless of how the code itself is divided up into different scripts in Ancillary-file-science, a typical ANTS based ancillary generation pipeline will usually consist of the following key stages:
Load Master File#
To get started generating an ancillary file, the first thing you will need to do is load a source dataset, referred to as “Master File”. Ideally this will be a netCDF file, but typically anything Iris can load can be loaded at this point. Other source formats may be supported via either specific ANTS routines or by other python libraries present in the environment.
Source Grid Processing#
If the source data is not already in the form of an Iris cube at this point, you will want to write a routine to convert it to one.
With the source data loaded in, at this point any source specific processing can be carried out. This might be metadata correction, selecting some subset of the source, or (more generally) “fixing” any problems with the source data. Typically this will take the form of Python code using Iris operations to work with data stored in Iris cubes.
In the case where the source grid processing is carried out in the pipeline as a distinct script, the application implementing that script is sometimes referred to as a “pre-processor”.
Regrid#
Once any source grid specific processing has been carried out, the next step is
to regrid the data to the target model grid. This can be carried out either by
using the standalone ants.cli.ancil_general_regrid application or calling the
appropriate ants.regrid routine.
Model Grid Processing#
With the data now on the model grid, at the target resolution, any grid specific operations can now be carried out. These might be filtering operations, generating derivative data fields, mappings, etc. This will take the form of Python code operating on Iris cubes. This code might use generic operations available via the ANTS library or more specific science routines implemented in Ancillary-file-science. At the end of this step, all science operations should have been applied, required metadata set, and all required data generated ready for saving.
Save to Ancillary File Format#
The final step is to save the processed data to the ancillary file format
appropriate for the model you are running, using the relevant saver routine
from the ants.io.save module.
Recommendations#
When writing the code going into your ancillary generation pipeline we make the following recommendations:
Work with netCDF files throughout the pipeline up until the final saving of the model ancillary file.
Do check the API docs to see if there is an existing ANTS routine that will carry out the operations you need.
Do try and split your code up into the distinct steps above where possible - monolithic scripts covering multiple steps can be hard to debug and expensive to keep re-running should problems occur during the execution of the script.
While the decomposition framework can be useful for supporting scalability of operations in a pipeline, it should be used sparingly. Only use it where you need to!